The Antibiotic
Resistance
Crisis
Antibiotic resistance is an escalating global crisis, where the spread of drug resistance is outpacing the development of new antibiotics.
Infections due to antibiotic-resistant bacteria are more dangerous for patients and more difficult and expensive to treat. They often require extended hospital stays, additional follow-up doctor visits, and more costly and toxic alternative treatments.1
As drug resistant bacteria become more common, so will deaths from life-threatening infections like sepsis, which is the body’s unregulated immune response to a severe bloodstream infection. Antibiotic resistance already causes at least 700,000 deaths a year and is expected to cause over 10 million deaths annually by 2050.2
Sepsis: Leading Cause of Death and Most Expensive Hospital Treated Condition in U.S. Hospitals.3-8
- Affects 1.7M adults per year
- Primary cause of 1 in 3 hospital deaths in the US
- In 2013, sepsis was estimated to cost the US $24B in hospital expenses
- Patients diagnosed with sepsis are 8x more likely to die during hospitalization
Early and effective treatment of antibiotic resistant infections has been proven to save lives. However, current technologies cannot provide actionable information quickly enough to enable early, targeted antibiotic treatment.
Day Zero is developing desperately needed diagnostics that rapidly identity pathogens and determine which antibiotics they will be most vulnerable to. This will allow doctors to provide the most effective treatment immediately, putting the brakes on life-threatening infections and saving countless lives.
Current Treatment
Minimizing the Time to Effective Antibiotic
Administration is Critical
Antibiotics are critical tools for fighting life-threatening bacterial infections, but the speed with which the patient is provided the appropriate antibiotic can be the difference between life and death. For example, the risk of death from septic shock increases by almost 8% for each hour an infection goes without appropriate treatment.9
Risk of Mortality, Septic Shock
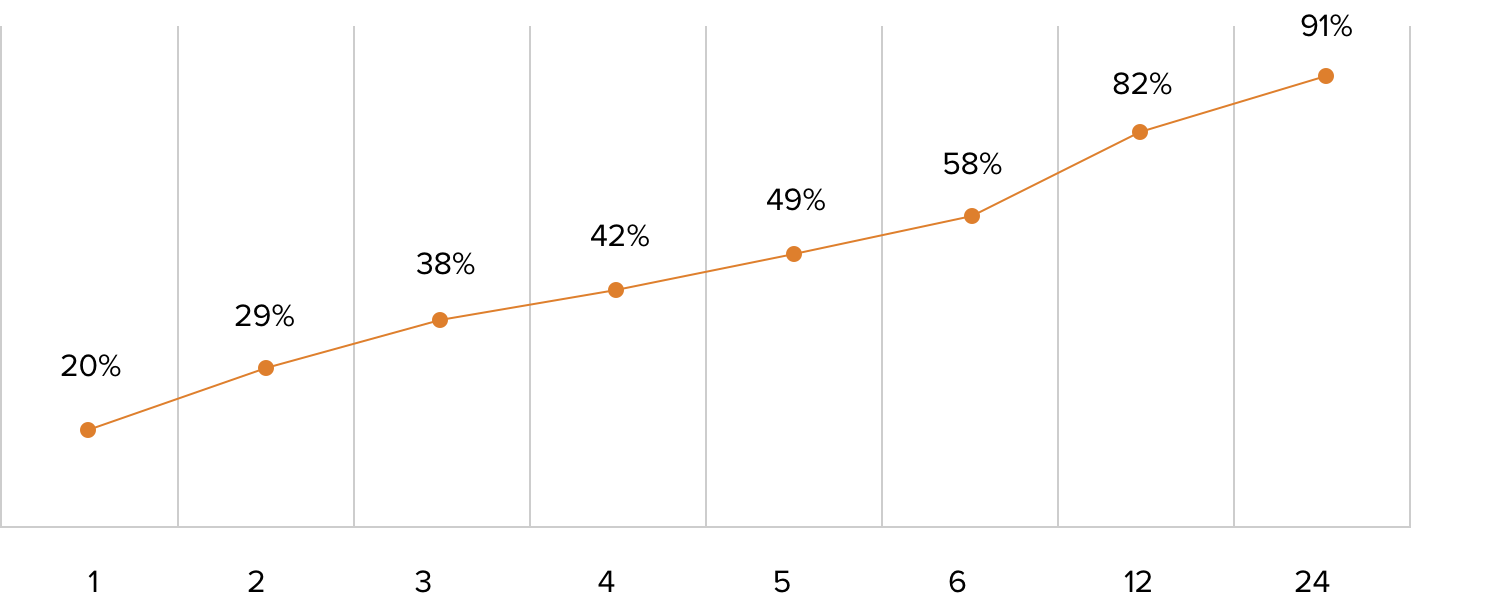
Hours to Treatment
When a patient shows up at a hospital today with signs of an infection, samples are collected and sent to the lab, where they are cultured on various media to see if they grow. It is only after the bacteria are allowed to grow for 1-2 days that they can be identified and tested for drug resistance. This antiquated process takes a total of 2-5 days and fails in 40% of severe infections. In the meantime, every hour that a patient remains inadequately treated increases their risk of complications or death.10
With little information to go on, physicians are forced to treat with empiric therapy: powerful, broad-spectrum antibiotics that are expensive, can have significant toxicity, and are increasingly less effective due to the spread of multidrug resistant pathogens. This practice also leads to higher rates of organ injury, increased risk of Clostridium difficile infection, and contributes to the growth of antibiotic resistance.11-13
Without innovative diagnostic approaches that bypass the need for time-consuming cultures, the world’s ability to prevent deaths from bacterial infections and address the growing threat of antibiotic resistance will be severely limited.
Day Zero’s
Solution
Day Zero is pioneering a new class of infectious disease diagnostics using whole-genome sequencing and machine learning to revolutionize how the world fights the growing threat of antibiotic resistance.
We are developing a diagnostic system that is intended to help physicians quickly and accurately diagnose and treat life-threatening infections without the need for a culture.
The system promises to help patients with severe infections receive the most effective antibiotic treatment on the first day they are admitted to the hospital — day zero.
References
- Centers for Disease Control and Prevention.
- IACG Report on Antimicrobial Resistance, April 2019.
- Rhee C et al. JAMA 2017; 318(13):1241-1249.
- Paoli C et al. Critical Care Medicine 2018; 46(12):1889–1897.
- Rhee C et al. JAMA Network Open 2019; 2(2):e187571.
- US Mortality Report 2017.
- Hall M et al. NCHS Data Brief 2011; 62.
- Torio CM et al. AHRQ Statistical Brief 2013; 204.
- Kumar A et al. Critical Care Med 2006; 34(6):1589-96.
- de Prost et al. Critical Care 2013; 17(5):1001.
- Singer M et al. Am J Respir and Crit Care Med 2017; 196(7).
- Jiang M et al. Front Cell Neurosci 2017; 11(308).
- Rhee C et al. JAMA Network Open 2020; 3(4):e202899.