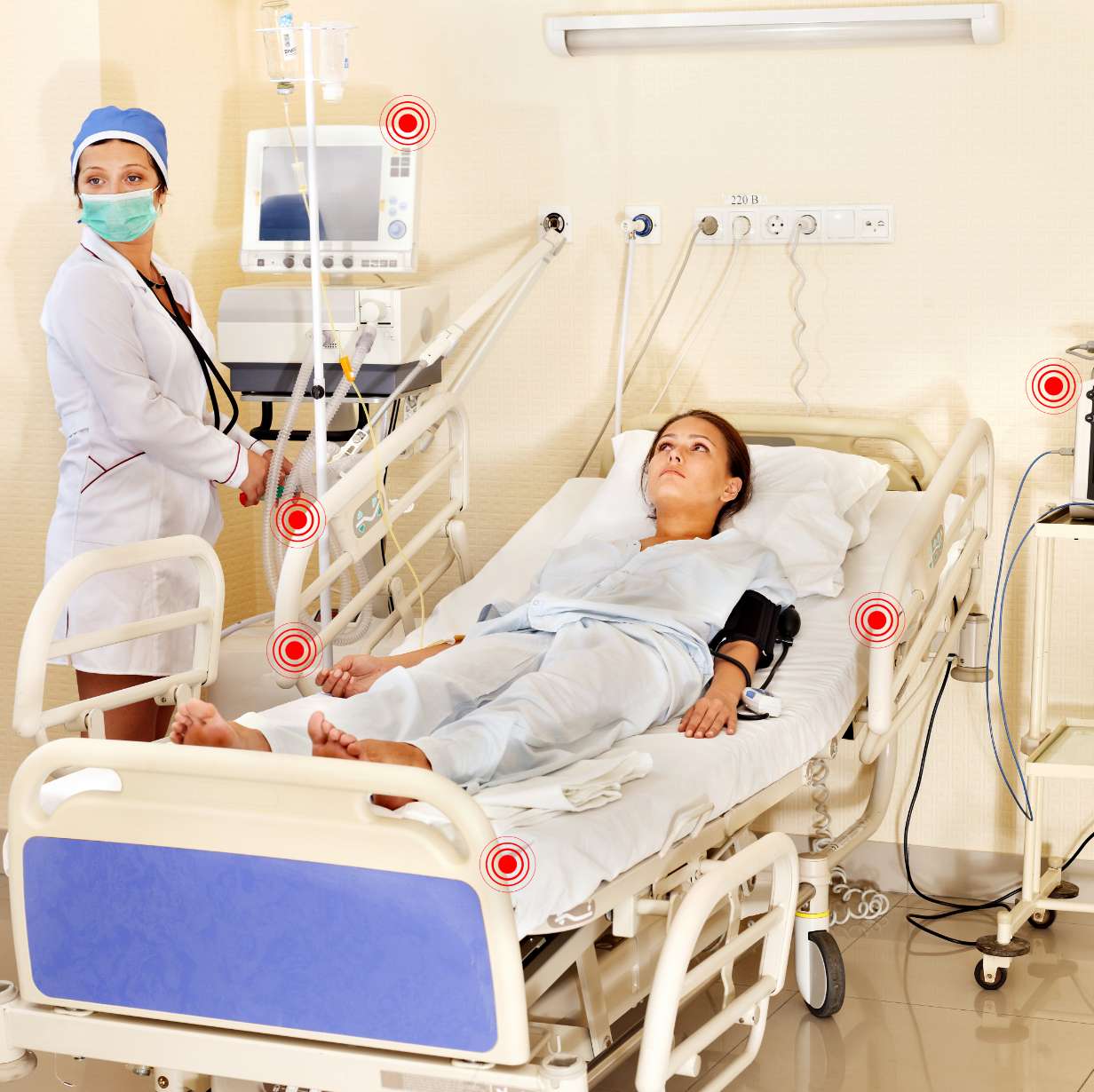
HAIs THREATEN PATIENT SAFETY
Healthcare-associated infections (HAIs) are a widespread challenge for patient care, annually affecting 4% of hospitalized patients in the U.S.1 and resulting in 99,000 patient deaths per year.2
The rising prevalence of multi-drug resistant organisms (MDROs) has made HAIs particularly dangerous, increasing the pressure for rapid, effective interventions when an outbreak is suspected.3,4